Article
In Silico Evaluation of Deleterious SNPs in Chicken TLR3 and TLR4 Genes
Donghyun Shin1, Ki-Duk Song2,3,†
Author Information & Copyright ▼
1Research Professor, Department of Animal Biotechnology, Chonbuk National University, Jeonju 54896, Republic of Korea
2Professor, Department of Animal Biotechnology, Chonbuk National University, Jeonju 54896, Republic of Korea
3Professor, The Animal Molecular Genetics and Breeding Center, Chonbuk National University, Jeonju 54896, Republic of Korea
ⓒ Copyright 2018, Korean Society of Poultry Science. This is an Open-Access article distributed under the terms of the
Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits
unrestricted non-commercial use, distribution, and reproduction in any
medium, provided the original work is properly cited.
Received: Jul 23, 2018 ; Revised: Sep 7, 2018 ; Accepted: Sep 07, 2018
Published Online: Sep 30, 2018
ABSTRACT
The innate immune recognition is based on the detection of microbial products. Toll-like receptors (TLRs) located on the cell surface and the endosome senses microbial components and nucleic acids, respectively. Chicken TLRs mediate immune responses by sensing ligands from pathogens, have been studied as immune adjuvants to increase the efficacy of vaccines. Single nucleotide polymorphisms (SNPs) of TLR3 and TLR4 genes in chicken were associated with resistance and susceptibility to viral infection. In this study, SNPs of chTLR3 and chTLR4 genes were retrieved from public database and annotated with chicken reference genome. Three-dimensional models of the chTLR3 and chTLR4 proteins were built using a Swiss modeler. We identified 35 and 13 nsSNPs in chTLR3 and chTLR4 genes respectively. Sorting Intolerant from Tolerant (SIFT) and Polymorphism Phenotyping v2 (Polyphen-2) analyses, suggested that, out of 35 and 13 nsSNPs, 4 and 2 SNPs were identified to be deleterious in chTLR3 and chTLR4 gene respectively. In chTLR3, 1 deleterious SNP was located in ectodomain and 3 were located in the Toll / IL-1 receptor (TIR) domain. Further structural model of chTLR3-TIR domain suggested that 1 deleterious SNP be present in the B-B loop region, which is important for TIR-TIR domain interactions in the downstream signaling. In chTLR4, the deleterious SNPs were located both in the ectodomain and TIR domain. SNPs predicted for chTLR3 and chTLR4 in this study, might be related to resistance or susceptible to viral infection in chickens. Results from this study will be useful to develop the effective measures in chicken against infectious diseases.
Keywords: chicken; polyphen-2; protein structure; SIFT; single nucleotide polymorphism; toll like receptors
INTRODUCTION
Microbial pathogens can be controlled by innate or adaptive immune systems. Detection of foreign molecules by innate immune cells is the first response to them, and it relies on sensing the constitutive and preserved products of microbial metabolism, whereas adaptive immune cells detects foreign antigens using their cognate receptors on the cell surface, i.e., T cell receptors (TCRs) and B cell receptors (BCRs) in animal (Medzhitov, 2001). Recognition of the conserved characteristics of microbial pathogens by the innate immune cells is mediated by pattern recognition receptors (PRRs). PRR detects conserved pathogen-associated molecular patterns (PAMPs) such as bacterial and viral cell wall components as well as nucleic acids (Iwasaki and Medzhitov, 2015). Toll-like receptors (TLRs) are crucial PRRs, localized on the cell surface as well as in the endosomes. TLRs are composed of an extracellular domain (ED) with leucine-rich repeats (LRRs) that recognize PAMPs, the transmembrane domain (TMD) and the cytosolic Toll / IL-1 receptor (TIR) domain, which allows binding of adaptor proteins, thereby initiates downstream signaling (Botos et al., 2011). Studies of TLRs have revealed pathogen detection mechanisms that induce the induction of cytokines and type I interferons (IFNs). Extracellular TLRs recognize the glycoprotein from the surface of the virus capsid, whereas endosomal TLRs senses the viral nucleic acids, such as dsRNA (O'neill et al., 2013).
The immunostimulatory properties of TLR ligands have been studied in chickens and several TLRs have been reported to mediate responses to pathogens (Paul et al., 2013). TLR ligands have been studied and used as adjuvants to increase vaccine efficacy in chickens (St. Paul et al., 2014a; St. Paul et al., 2014b). Previous studies identified 10 TLRs (TLR1LA, ILB, 2A, 2B, 3, 4, 5, 7, 15 and 21) in chickens (Kalaiyarasu et al., 2016; Rahim et al., 2017). Among chicken TLRs, TLR3 (chTLR3) and TLR4 (chTLR4) that are well studied receptors, recognize double-stranded RNA (dsRNA) and lipopolysaccharide (LPS), respectively. TLRs are polymorphic among different chicken breed that may be related to resistance or susceptibility to viral infections (Philbin et al., 2005; Ruan et al., 2012; Ruan et al., 2015).
The most common genetic variation in the genome is single nucleotide polymorphisms (SNPs) and the frequency of distribution is estimated to be 1 every 300-1,000 bp (Kruglyak and Nickerson, 2001). SNPs represent unique genetic markers and important variations that can induce phenotypic changes as well as causing diseases (Shastry, 2009). In particular, nonsynonymous SNPs (nsSNPs) that alter the amino acid sequence have been extensively studied for their ability to be readily detected and to alter protein function (Wang and Sadée, 2006).
In this study, we have identified and annotated SNPs of chTLR3 and chTLR4 genes from Ensembl database. In addition, we predicted the deleterious SNPs using computational tools such as the Sorting Intolerant from Tolerant (SIFT) and Polymorphism Phenotyping v2 (Polyphen-2). Finally, we have modelled the three-dimensional structures of chTLR3 and chTLR4 and mapped the location of the deleterious SNPs. This study may provide some insight on the functional impact of SNPs in chTLR3 and chTLR4 genes, which may be useful for selective breeding as well as for studying mechanisms of disease resistance or susceptibility in chickens.
MATERIALS AND METHODS
1. SNP Data Retrieval and Prediction of Deleterious SNPs in Chicken TLR3 and TLR4 Genes
For this study, we retrieved SNPs of chTLR3 and chTLR4 gene from Ensembl database (http://useast.ensembl.org/Gallus_ gallus/Info/Index) using available variant calling file (VCF) for chicken genome (Gallus_gallus-5.0.86.; http://useast.ensembl. org/Gallus_gallus/Info/Index?db=core). Further nsSNP-based sequence homology tests were performed to identify important amino acid substitutions that might affect biological functions via structural modifications of proteins. SIFT uses sequence homology to predict the effects of the amino acid substitution of interest on protein functions (Ng and Henikoff, 2003). In our analysis 35 nsSNPs from chTLR3 gene and 13 nsSNPs from chTLR4 gene were identified from SIFT analysis with cut-off score <0.05. In addition, we performed PolyPhen-2 analysis on the SIFT results in order to identify the deleterious nsSNPs. Tested nsSNPs were classified as probably damaging, possibly damaging, or benign if they received Polyphen-2 scores (range: 0 to 1) of >0.95, 0.5 to 0.95, or <0.5, respectively (Adzhubei et al., 2013). In this study, we expected that probably damaging and possibly damaging SNPs would affect protein functions.
2. Structural Modelling of chTLR3 and chTLR4
We have built 3-D structural models for chTLR3 and chTLR4 using SWISS-MODEL (Waterhouse et al., 2018). At first initial sequence has been extracted from the UniProt (UniProt ID: F1NG27) and we analyzed the domains using SMART domain database (http://smart.embl-heidelberg.de/). We have considered only ectodomain and TIR domain for both the chTLR3 and chTLR4. HumanTLR4 (4G8A) and mouse TLR3 ectodomain (3CIY) structures were used as templates to construct ectodomain of chTLR3 and chTLR4. To construct TIR domain model we have used TLR10 TIR-domain (2J67) as a template for both chTLR3 and chTLR4. Energy minimization using steepest descent and conjugate gradient was performed on the models using chimera. Since we have not observed deleterious mutations in the ECD, we mainly focused on the TIR domain and the dimerization. We performed modeling for mutations using Discovery Studio Visualizer by replacing original amino acid with proper orientation for the side chain.
RESULTS AND DISCUSSION
1. Classification of SNPs Present in the chTLR3 and chTLR4 Gene
Variants present in the chTLR3 and chTLR4 genes were classified according to location and function. Total of 402 and 109 variants were observed in the chTLR3 and chTLR4 respectively (Table 1). However, we observed only few nsSNPs in the coding region of both TLRs; 35 nsSNPs (8.71% of the total SNPs) for chTLR3 and 13 nsSNPs (11.93% of the total SNPs) for chTLR4 were identified (Table 2 and 3). Moreover, we observed large number of SNPs at the intronic region of both TLRs; 344 (85.57% of the total SNPs) and 54 SNPs (49.54% of the total SNPs) for chTLR3 and chTLR4 respectively (Table 1).
Table 1.
Count of SNPs variant in Chicken TLR3 and TLR4 genes
| Variant |
chTLR3
|
chTLR4
|
Total |
| 3 prime UTR-variant |
|
23 |
23 |
| Frameshift-variant |
1 |
|
1 |
| Intronic-variant |
344 |
54 |
398 |
| Missense-variant |
35 |
13 |
48 |
| Splice-donor-variant |
1 |
|
1 |
| Splice-region-variant |
3 |
1 |
4 |
| Stop-gained-variant |
|
1 |
1 |
| Synonymous-variant |
18 |
17 |
35 |
Download Excel Table
Table 2.
The prediction of nsSNPs influence on protein function with Polyphen-2 and SIFT in chicken TLR3 (The numbers given in the bracket is amino acid number used from UniProt sequence ID: Q0PQ88)
| #Uploaded-variation |
Location |
REF |
ALT |
IMPACT |
Feature type |
EXON |
Protein position |
Amino acids |
SIFT |
SIFTscore |
Polyphen |
Polyphen 2 score |
| rs80605562 |
4:61707236-61707236 |
A |
C |
MODERATE |
Transcript |
1/9 |
12 |
Q/P |
deleterious_low_confidence |
0.02 |
possibly damaging |
0.722 |
| rs80692549 |
4:61707399-61707399 |
A |
T |
MODERATE |
Transcript |
2/9 |
19(68) |
D/V |
tolerated |
0.47 |
benign |
0.009 |
| rs14479722 |
4:61707470-61707470 |
A |
C |
MODERATE |
Transcript |
2/9 |
43(92) |
K/Q |
tolerated |
0.29 |
benign |
0.005 |
| rs14479723 |
4:61707489-61707489 |
A |
G |
MODERATE |
Transcript |
2/9 |
49(98) |
N/S |
tolerated |
0.52 |
benign |
0.003 |
| rs731047275 |
4:61707585-61707585 |
A |
G |
MODERATE |
Transcript |
2/9 |
81(130) |
N/S |
tolerated |
0.15 |
benign |
0.247 |
| rs80658674 |
4:61708591-61708591 |
A |
G |
MODERATE |
Transcript |
3/9 |
131(180) |
S/G |
tolerated |
0.45 |
benign |
0.002 |
| rs80619479 |
4:61709615-61709615 |
A |
C |
MODERATE |
Transcript |
4/9 |
174(232) |
N/H |
tolerated |
0.11 |
benign |
0.421 |
| rs80733702 |
4:61709628-61709628 |
G |
A |
MODERATE |
Transcript |
4/9 |
178(236) |
S/N |
tolerated |
0.5 |
benign |
0.005 |
| rs80572088 |
4:61709896-61709896 |
A |
T |
MODERATE |
Transcript |
6/9 |
201(334) |
T/S |
tolerated |
0.75 |
benign |
0.023 |
| rs80713467 |
4:61710093-61710093 |
G |
C |
MODERATE |
Transcript |
6/9 |
266(401) |
R/S |
tolerated |
0.52 |
benign |
0.005 |
| rs80718952 |
4:61710143-61710143 |
A |
G |
MODERATE |
Transcript |
6/9 |
283(418) |
E/G |
tolerated |
1 |
benign |
0.001 |
| rs314125162 |
4:61710434-61710434 |
G |
A |
MODERATE |
Transcript |
6/9 |
380(515) |
R/K |
tolerated |
1 |
benign |
0 |
| rs794518640 |
4:61710634-61710634 |
A |
G |
MODERATE |
Transcript |
6/9 |
447(582) |
K/E |
tolerated |
0.16 |
possibly damaging |
0.63 |
| rs80778047 |
4:61710677-61710677 |
C |
T |
MODERATE |
Transcript |
6/9 |
461(596) |
A/V |
tolerated |
0.36 |
benign |
0.043 |
| rs80668052 |
4:61710691-61710691 |
G |
C |
MODERATE |
Transcript |
6/9 |
466(601) |
D/H |
tolerated |
0.13 |
benign |
0.051 |
| rs80636705 |
4:61710832-61710832 |
G |
T |
MODERATE |
Transcript |
6/9 |
513(648) |
A/S |
tolerated |
0.24 |
benign |
0.008 |
| rs80641674 |
4:61711004-61711004 |
C |
T |
MODERATE |
Transcript |
6/9 |
570(705) |
A/V |
tolerated |
0.41 |
benign |
0.002 |
| rs80747649 |
4:61711195-61711195 |
A |
T |
MODERATE |
Transcript |
6/9 |
634(769) |
T/S |
tolerated |
0.27 |
benign |
0.001 |
| rs80669462 |
4:61711252-61711252 |
G |
A |
MODERATE |
Transcript |
6/9 |
653(788) |
E/K |
tolerated |
0.2 |
possibly damaging |
0.894 |
| rs80661166 |
4:61711293-61711293 |
A |
G |
MODERATE |
Transcript |
6/9 |
666(801) |
I/M |
tolerated |
0.23 |
probably damaging |
0.995 |
| rs80757506 |
4:61712065-61712065 |
G |
A |
MODERATE |
Transcript |
7/9 |
716(851) |
D/N |
tolerated |
0.09 |
possibly damaging |
0.784 |
| rs735041008 |
4:61715833-61715833 |
C |
T |
MODERATE |
Transcript |
8/9 |
763 |
P/L |
deleterious_low_confidence |
0.04 |
unknown |
? |
| rs740030255 |
4:61715851-61715851 |
T |
G |
MODERATE |
Transcript |
8/9 |
769 |
V/G |
deleterious_low_confidence |
0 |
unknown |
? |
| rs316043187 |
4:61715914-61715914 |
A |
G |
MODERATE |
Transcript |
8/9 |
790 |
N/S |
deleterious_low_confidence |
0 |
unknown |
? |
| rs736982671 |
4:61715944-61715944 |
A |
G |
MODERATE |
Transcript |
8/9 |
800 |
Q/R |
deleterious_low_confidence |
0 |
unknown |
? |
| rs733651632 |
4:61715958-61715958 |
T |
G |
MODERATE |
Transcript |
8/9 |
805 |
S/A |
deleterious_low_confidence |
0 |
unknown |
? |
| rs316084677 |
4:61716029-61716029 |
G |
T |
MODERATE |
Transcript |
8/9 |
828 |
R/S |
deleterious_low_confidence |
0 |
unknown |
? |
| rs14479729 |
4:61720359-61720359 |
C |
T |
MODERATE |
Transcript |
9/9 |
830 |
P/L |
deleterious_low_confidence |
0 |
unknown |
? |
| rs14479730 |
4:61720422-61720422 |
C |
T |
MODERATE |
Transcript |
9/9 |
851 |
A/V |
deleterious_low_confidence |
0 |
unknown |
? |
| rs14479732 |
4:61720444-61720444 |
T |
A |
MODERATE |
Transcript |
9/9 |
858 |
H/Q |
deleterious_low_confidence |
0 |
unknown |
? |
| rs793896212 |
4:61720447-61720447 |
A |
C |
MODERATE |
Transcript |
9/9 |
859 |
Q/H |
deleterious_low_confidence |
0 |
unknown |
? |
| rs318110846 |
4:61720512-61720512 |
A |
G |
MODERATE |
Transcript |
9/9 |
881 |
D/G |
deleterious_low_confidence |
0 |
unknown |
? |
| rs1057909073 |
4:61720562-61720562 |
C |
T |
MODERATE |
Transcript |
9/9 |
898 |
P/S |
deleterious_low_confidence |
0 |
unknown |
? |
| rs1057733301 |
4:61720567-61720567 |
C |
G |
MODERATE |
Transcript |
9/9 |
899 |
C/W |
deleterious_low_confidence |
0 |
unknown |
? |
| rs738876733 |
4:61720632-61720632 |
G |
A |
MODERATE |
Transcript |
9/9 |
921 |
G/D |
deleterious_low_confidence |
0 |
unknown |
? |
Download Excel Table
Table 3.
The prediction of nsSNPs influence on protein function with polyphen and SIFT in chicken TLR4
| #Uploaded-variation |
Location |
REF |
ALT |
IMPACT |
Feature type |
EXON |
Protein position |
Amino acids |
SIFT |
SIFT_score |
Polyphen |
Polyphen 2 score |
| rs738395510 |
17:4083135-4083135 |
A |
C |
MODERATE |
Transcript |
1/3 |
8 |
T/P |
tolerated |
0.35 |
benign |
0 |
| rs739102600 |
17:4083154-4083154 |
T |
G |
MODERATE |
Transcript |
1/3 |
14 |
V/G |
tolerated |
0.21 |
benign |
0.233 |
| rs1058797423 |
17:4083181-4083181 |
C |
G |
MODERATE |
Transcript |
1/3 |
23 |
S/C |
deleterious |
0.04 |
probably damaging |
0.997 |
| rs1059739146 |
17:4083190-4083190 |
C |
T |
MODERATE |
Transcript |
1/3 |
26 |
A/V |
tolerated |
0.1 |
probably damaging |
0.991 |
| rs14102161 |
17:4084296-4084296 |
A |
G |
MODERATE |
Transcript |
2/3 |
83 |
K/E |
tolerated |
1 |
benign |
0 |
| rs732610021 |
17:4085707-4085707 |
G |
A |
MODERATE |
Transcript |
3/3 |
225 |
G/E |
deleterious |
0.01 |
probably damaging |
0.996 |
| rs312339277 |
17:4085815-4085815 |
G |
A |
MODERATE |
Transcript |
3/3 |
261 |
R/K |
tolerated |
0.25 |
benign |
0.121 |
| rs15032443 |
17:4085936-4085936 |
T |
G |
MODERATE |
Transcript |
3/3 |
301 |
D/E |
tolerated |
1 |
benign |
0 |
| rs15032441 |
17:4086061-4086061 |
A |
G |
MODERATE |
Transcript |
3/3 |
343 |
K/R |
tolerated |
0.41 |
benign |
0.027 |
| rs15032440 |
17:4086180-4086180 |
C |
T |
MODERATE |
Transcript |
3/3 |
383 |
H/Y |
tolerated |
0.25 |
benign |
0 |
| rs731654157 |
17:4086779-4086779 |
G |
T |
MODERATE |
Transcript |
3/3 |
582 |
L/F |
tolerated |
0.72 |
benign |
0.003 |
| rs15032438 |
17:4086865-4086865 |
G |
A |
MODERATE |
Transcript |
3/3 |
611 |
R/Q |
tolerated |
0.43 |
benign |
0 |
| rs735198923 |
17:4087413-4087413 |
A |
G |
MODERATE |
Transcript |
3/3 |
794 |
R/G |
deleterious |
0 |
possibly damaging |
0.805 |
Download Excel Table
2. Evaluation of Impact of nsSNPs in chTLR3 and chTLR4
SIFT and Polyphen-2 tools were used to predict the effect of nsSNPs in chTLR3 and chTLR4. For chTLR3, SIFT analysis predicted that 15 nsSNPs were deleterious (with low confidence score) and 20 nsSNPs were tolerated. In addition, Polyphen-2 analysis predicted 16 nsSNPs as benign, 4 nsSNPs as probable damaging, 1 nsSNP as possibly damaging and 14 nsSNPs as unknown (Table 2). For chTLR4, SIFT analysis predicted that 3 were deleterious and 10 were tolerated. Furthermore, Polyphen-2 analysis showed that 1 possibly damaging, 3 probably damaging and 9 nsSNPs as benign (Table 3). We selected total 8 nsSNPs from both chTLR3 and chTLR4 genes to map on the 3D structure of chTLR3 and chTLR4 to check the location of these damaging SNPs in the protein.
3. Mapping of Deleterious nsSNPs on the chTLR3 and chTLR4 Structures
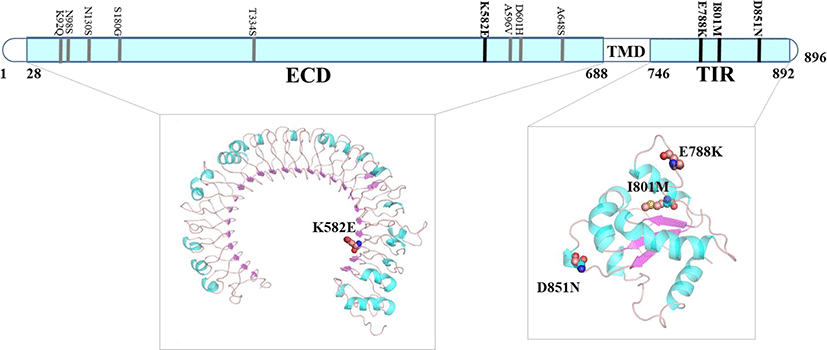
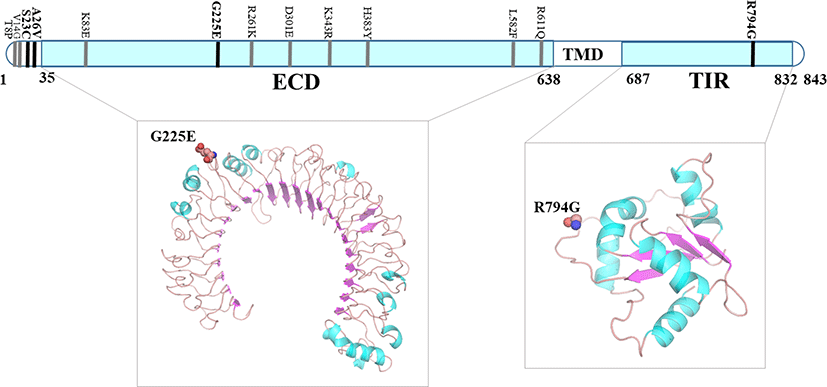
We have used UniProt sequence number for amino acids particularly for chTLR3 due to variation between chTLR3 transcript from Ensembl and chTLR3 protein sequence from UniProt database (UniProt ID: Q0PQ88). We performed comparative modeling in order to construct 3-dimensional models for chTLR3 and chTLR4. ECD of chTLR3 and chTLR4 shows typical concave surface and with 23 and 27 leucine rich repeats respectively. Protein structure analysis was performed using four nsSNPs sequence regions predicted to be deleterious among the chTLR3 nsSNPs, i.e., K582E, E788K, I801M, D851N. Among them, K582E amino acid variant was located at the ECD, whereas E788K, I801M and D851N were located in the TIR domain (Fig. 1). We have not identified deleterious SNPs on the core of the structure where ligand binds. Previous studies also have reported that there are six amino acid variants (D14V, R345S, G362E, R459K, A540V, and A649V) externally in the ECD of chTLR3 (Ruan et al., 2015). ChTLR4 is known to be involved in the recognition of bacterial components and Salmonella in innate immune responses. In this study, we identified 4 nsSNPs (S23C, A26V, G225E and R794G) for chTLR4 (Fig. 2). However, 2 of them S23C, A26V, were observed in the signal sequence (Fig 2). The five amino acid variants of chTLR4 (G225E, D301E, R343K, H383Y and R611Q) have been reported to be associated with chicken susceptibility to salmonellosis (Leveque et al., 2003). Six amino acid variants (K83E, R261K, F427V, P551T, E547D and K714R), which are thought to be related to susceptibility and resistance to Salmonella infection in chicken, have been reported (Ruan et al., 2012).
Fig. 1.
nsSNPs predicted to affect protein structure in the chTLR3 ECD and TIR (ECD: K582E, TIR: E788K, I801M, D851N are predicted to be deleterious).
Download Original Figure
Fig. 2.
nsSNPs predicted to affect protein structure in the TLR4 ECD and TIR (ECD: G225E, TIR: R794G are predicted to be deleterious).
Download Original Figure
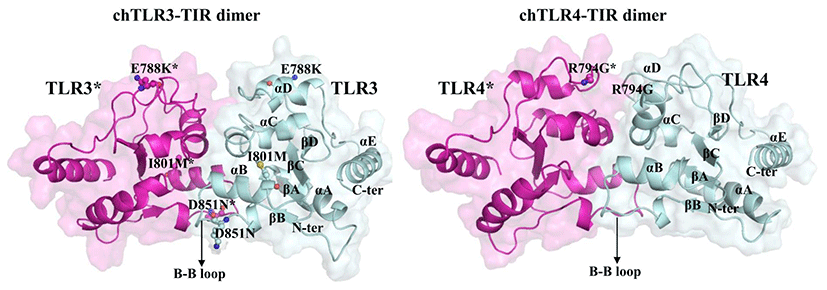
We further focused on the TIR domain of the chTLR3 where intracellular adaptors will form complexes required for signaling cascades. chTLR3-TIR domain shows one mutation presents in the BB-loop region which is crucial for the dimerization of the adaptor proteins in TLR signaling (Fig. 3), whereas other deleterious mutations located at the other loop regions. In chTLR4 TIR domain, only one deleterious mutation was identified and located in the alpha-C region (Fig. 3). Hence these mutations may have an impact on the homo-dimerization as well as affect the dimerization and recruitment of adaptor proteins for the downstream signaling, thereby leads to cumulative effect on the whole signaling. TLR3 is involved in the recognition of viral dsRNA in chickens. ChTLR3 expression in chickens is associated with Newcastle disease virus and avian influenza virus infection (Stewart et al., 2012; Cheng et al., 2014). In addition, polymorphisms of the chTLR3 gene have been reported to be associated with susceptibility to various diseases such as hepatitis B virus infection and chronic hepatitis C (Al-Qahtani et al., 2012; Lee et al., 2013). Thus, it is reasonable to assume that the polymorphic sites found in the chTLR3 gene may be involved in susceptibility or susceptibility to chicken viral infection. Moreover, we have constructed dimer model of chTLR3-TIR domain.
Fig. 3.
Visualization of the predicted deleterious nsSNPs on the TIR domain dimer. In particular, D851N locates in the BB-loop of chTLR3.
Download Original Figure
In this study, we have classified polymorphic sites in the chTLR3 and chTLR4 genes. We identified 402 SNPs and 109 SNPs in the chTLR3 and chTLR4 genes, respectively. Similar to previous studies about chTLR3 and chTLR4, we have detected several amino acid variations in chickens. The mutations that could have impact the function in chTLR3 and chTLR4 were predicted and analyzed for protein structure. These SNPs may be associated with susceptibility and resistance to virus infection in chickens. This study may provide a basis for future research on viral diseases in chickens.
ACKNOWLEGEMENTS
This work was supported by the Next-Generation BioGreen 21 Program (No. PJ01324201, PJ01315101), Rural Development Administration, Republic of Korea, and the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries (IPET) through the Agriculture, Food and Rural Affairs Research Center Support Program, funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA; grant number 716002-7).
References
Adzhubei I, Jordan DM, Sunyaev SR 2013 Predicting functional effect of human missense mutations using PolyPhen 2. Current Protocols in Human Genetics 76(1):7.20.1- 7.20.41 .

Kalaiyarasu S, Bhatia S, Mishra N, Sood R, Kumar M, SenthilKumar D, Bhat S, Prakash MD 2016 Elevated level of pro inflammatory cytokine and chemokine expression in chicken bone marrow and monocyte derived dendritic cells following LPS induced maturation. Cytokine 85: 140-147 .

Rahim A, Kumar S, Debnath J, Yadav R, Das AK, Yadav A 2017 Fold changes in relative mRNA expression of immune response genes in lymphoid tissues of Rhode Island Red chicken. Indian Journal of Animal Sciences 87(6):108-110 .

Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T 2018 SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res 46(W1):W296-W303 .